- #Macvector aligning illumina reads to a reference sequence how to
- #Macvector aligning illumina reads to a reference sequence install
- #Macvector aligning illumina reads to a reference sequence software
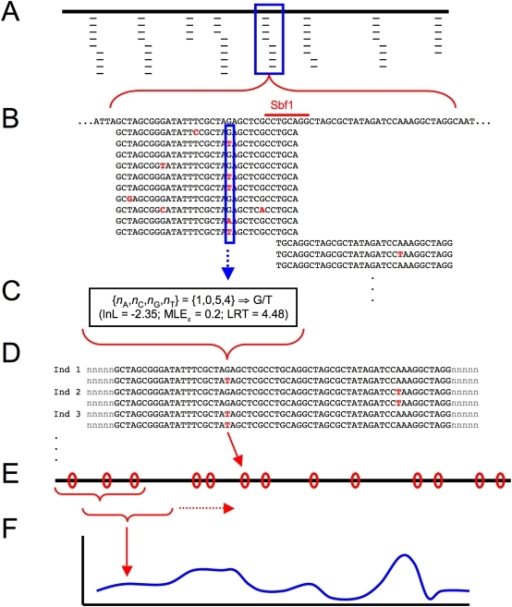
Details of the nucleotide alignments of the primer target regions of 271 nrfA sequences from reference genomes representing 18 distinct clades of NrfA are shown here along with validation of application to PCR-based methodology including the use of amplified fragment length polymorphism (AFLP) profiling and Illumina platform amplicon-based sequencing of environmental samples and selected reference strains. Trimmomatic output files will show which reads (if any) were trimmed.PCR primer sets were designed to target nrfA, the gene encoding the pentaheme nitrite reductase NrfA that catalyzes the nitrite ammonification step in the process of dissimilatory nitrate reduction to ammonium (DNRA). This file can contain multiple adapter sequences by using a multi-FASTA file format. The second record contains the adapter sequence. The first record contains the right-caret character followed by an arbitrary string. In our example, using the Nextera XT library prep kit, the “adapters.fasta” file would look like this:
#Macvector aligning illumina reads to a reference sequence how to
See the manual for more information about how to set these three fields. This parameter also requires three additional fields: seedMismatches, palindromeClipThreshold, simpleClipThreshold. The ILLUMINACLIP parameter specifies the name of this file. The adapter sequence(s) is/are contained in a FASTA formatted file.For each read file, we specify the name of a paired output file and an unpaired output file.We specify both files in the parameter list. There are always two FASTQ files in a paired-end run: one file for the forward reads and one file for the reverse reads.To speed up the application, we specify the number of threads to use, up to the maximum number of available processor threads.Most sequencing runs use paired-end reads, so we specify “PE” in the command line.If you decide to use Trimmomatic for trimming adapter sequences from Illumina reads, a minimal command that only performs adapter trimming may look like this: java jar trimmomatic-0.39.jar PE -threads 4 read1_ read1_ read2_ read2_ ILLUMINACLIP:adapters.fasta:2:30:10
#Macvector aligning illumina reads to a reference sequence install
The Trimmomatic manual describes how to install this application, how to run it and it describes all of the required and optional command line parameters. Trimmomatic is a popular tool for trimming adapter sequences from Illumina reads. If you wish to use any of these applications, both FASTQ and FASTA files are available for that purpose.
#Macvector aligning illumina reads to a reference sequence software
Adapter trimming software applications may require FASTQ or FASTA formatted files.
Our bioinformatics data analysis pipelines automatically convert FASTQ files to FASTA files, so you will receive both file formats at the end of each sequencing run. They represent the raw data files from which all downstream processing begins. FASTQ files are automatically generated by Illumina sequencers. We deliver sequencing results in FASTQ and FASTA file formats. The adapter sequences for other kits may be different, so be sure to check which kit was used for library prep and get the appropriate sequence from the adapter sequences manual. Illumina Nextera XT is one of our most used kits. The exact DNA sequence of the adapters depends on the library preparation kit that is used for sequencing. This step will improve downstream data processing, such as sequence alignments and de novo assembly. When sequencing is complete it’s important to remove or trim off, the adapter sequences from the reads. The adapter sequences are required for attaching reads to flow cells and for attaching indexes to reads. During the library preparation process, Illumina adapter sequences are annealed to sequencing reads.
